Crops ›› 2025, Vol. 41 ›› Issue (5): 135-141.doi: 10.16035/j.issn.1001-7283.2025.05.018
Previous Articles Next Articles
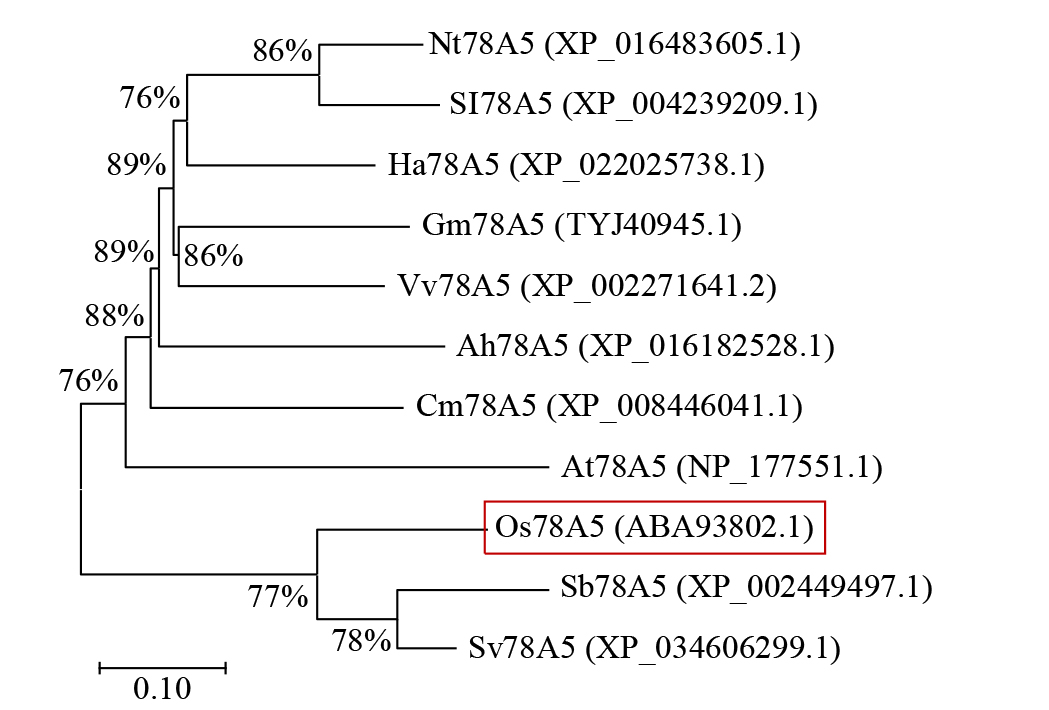
Expression Analysis of the CYP450 Family Gene Os78A5 in Rice
Zhi Xianhong1( ), Ji Zixian1, Xu Zhenwang1, Tan En1, Liang Rishen1, Ma Shuaipeng2, Tang Huiwu1(
), Ji Zixian1, Xu Zhenwang1, Tan En1, Liang Rishen1, Ma Shuaipeng2, Tang Huiwu1( )
)
- 1
College of Agriculture and Biology ,Zhongkai University of Agriculture and Engineering Guangzhou 510225, Guangdong, China
2Guangzhou Experimental Station ,Chinese Academy of Tropical Agricultural Sciences Guangzhou 510145, Guangdong, China
| [1] | 杨杰, 詹亚光, 肖佳雷, 等. 细胞色素 P450 在植物三萜和甾醇骨架修饰中的功能研究进展. 中国科学:生命科学, 2018, 48(10):1065-1083. |
| [2] |
Guengerich F P. Cytochrome P450 research and the journal of biological chemistry. Journal of Biological Chemistry, 2019, 294(5):1671-1680.
doi: 10.1074/jbc.TM118.004144 pmid: 29871932 |
| [3] | Nelson D R. Cytochrome P450 diversity in the tree of life. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics, 2018, 1866(1):141-154. |
| [4] |
Nelson D, Werck‐Reichhart D. A P450‐centric view of plant evolution. The Plant Journal, 2011, 66(1):194-211.
doi: 10.1111/j.1365-313X.2011.04529.x pmid: 21443632 |
| [5] | Bak S, Beisson F, Bishop G, et al. Cytochromes P450. The Arabidopsis Book, 2011, 9:e0144. |
| [6] | 罗浑金, 傅童声. 细胞色素P450的多样性与介导抗性的研究. 生物技术通报, 2006(2):14-16. |
| [7] | Chapple C. Molecular-genetic analysis of plant cytochrome P450- dependent monooxygenases. Annual Review of Plant Biology, 1998, 49(1):311-343. |
| [8] | Wang Y F, Ahmad N, et al. Unraveling the functional characterization of a jasmonate-induced flavonoid biosynthetic CYP45082G24 gene in Carthamus tinctorius. Functional & Integrative Genomics, 2023, 23(2):172. |
| [9] | Wang H Y, Wang Y, et al. Genome-wide identification of the CYP82 gene family in cucumber and functional characterization of CsCYP82D102 in regulating resistance to powdery mildew.. PeerJ, 2024, 12:e17162. |
| [10] |
He J, Chen Q W, Xin P Y, et al. CYP72A enzymes catalyse 13-hydrolyzation of gibberellins. Nature Plants, 2019, 5(10):1057-1065.
doi: 10.1038/s41477-019-0511-z pmid: 31527846 |
| [11] | Pandian B A, Sathishraj R, Djanaguiraman M, et al. Role of cytochrome P450 enzymes in plant stress response. Antioxidants, 2020, 9(5):454. |
| [12] | Rao M J, Xu Y, Tang X, et al. CsCYT75B1, a citrus CYTOCHROME P 450 gene, is involved in accumulation of antioxidant flavonoids and induces drought tolerance in transgenic Arabidopsis. Antioxidants, 2020, 9(2):161. |
| [13] | Yan Q, Cui X X, Lin S, et al. GmCYP82A3,a soybean cytochrome P 450 family gene involved in the jasmonic acid and ethylene signaling pathway, enhances plant resistance to biotic and abiotic stresses. PLoS ONE, 2016, 11(9):e0162253. |
| [14] |
Jiang L, Yoshida T, Stiegert S, et al. Multi-omics approach reveals the contribution of KLU to leaf longevity and drought tolerance. Plant Physiology, 2021, 185(2):352-368.
doi: 10.1093/plphys/kiaa034 pmid: 33721894 |
| [15] | 李泽宇.OsCYP71D8L 调控水稻株高和耐逆性的分子基础. 北京: 中国农业科学院, 2021. |
| [16] | Kurotani K, Hayashi K, Hatanaka S, et al. Elevated levels of CYP94 family gene expression alleviate the jasmonate response and enhance salt tolerance in rice. Plant and Cell Physiology, 2015, 56(4):779-789. |
| [17] | Maeda S, Dubouzet J G, Kondou Y, et al. The rice CYP78A gene BSR2 confers resistance to Rhizoctonia solani and affects seed size and growth in Arabidopsis and rice. Scientific Reports, 2019, 9(1):587. |
| [18] | Wang A J, Ma L, Shu X Y, et al. Rice (Oryza sativa L.) cytochrome P 450 protein 716A subfamily CYP716A16 regulates disease resistance. BMC Genomics, 2022, 23(1):343. |
| [19] |
Sakamoto T, Kawabe A, Tokida‐Segawa A, et al. Rice CYP734As function as multisubstrate and multifunctional enzymes in brassinosteroid catabolism. The Plant Journal, 2011, 67(1):1-12.
doi: 10.1111/j.1365-313X.2011.04567.x pmid: 21418356 |
| [20] | Zhou C L, Lin Q B, Ren Y L, et al. A CYP78As-small grain4-coat protein complex Ⅱ pathway promotes grain size in rice. The Plant Cell, 2023, 35(12):4325. |
| [21] | Sahoo B, Nayak I, Parameswaran C, et al. A comprehensive genome-wide investigation of the cytochrome 71 (OsCYP71) gene family: revealing the impact of promoter and gene variants (Ser33Leu) of OsCYP71P6 on yield-related traits in indica rice (Oryza sativa L.). Plants, 2023, 12(17):3035. |
| [22] | Song S F, Wang T K, et al. A novel strategy for creating a new system of third‐generation hybrid rice technology using a cytoplasmic sterility gene and a genic male‐sterile gene. Plant Biotechnology Journal, 2021, 19(2):251-260. |
| [23] | Li H, Pinot F, Sauveplane V, et al. Cytochrome P450 family member CYP704B2 catalyzes the ω-hydroxylation of fatty acids and is required for anther cutin biosynthesis and pollen exine formation in rice. The Plant Cell, 2010, 22(1):173-190. |
| [24] | Huang X Y, Chang Z Y, et al. Regulation by distinct MYB transcription factors defines the roles of OsCYP86A9 in anther development and root suberin deposition. The Plant Journal, 2024, 118(6):1972-1990. |
| [25] | Cui Y J, Peng Y L, Zhang Q, et al. Disruption of EARLY LESION LEAF 1, encoding a cytochrome P 450 monooxygenase, induces ROS accumulation and cell death in rice. The Plant Journal, 2021, 105(4):942-956. |
| [26] | Berens M L, Wolinska K W, Spaepen S, et al. Balancing trade- offs between biotic and abiotic stress responses through leaf age- dependent variation in stress hormone cross-talk. Proceedings of the National Academy of Sciences of the United States of America, 2019, 116(6):2364-2373. |
| [27] |
Mafu S, Ding Y, Murphy K M, et al. Discovery, biosynthesis and stress-related accumulation of dolabradiene-derived defenses in maize. Plant Physiology, 2018, 176(4):2677-2690.
doi: 10.1104/pp.17.01351 pmid: 29475898 |
| [28] |
Xu J, Wang X Y, Guo W Z. The cytochrome P450 superfamily: Key players in plant development and defense. Journal of Integrative Agriculture, 2015, 14(9):1673-1686.
doi: 10.1016/S2095-3119(14)60980-1 |
| [29] |
Tao X, Wang M X, Dai Y, et al. Identification and expression profile of CYPome in perennial ryegrass and tall fescue in response to temperature stress. Frontiers in Plant Science, 2017, 8:1519.
doi: 10.3389/fpls.2017.01519 pmid: 29209335 |
| [30] | Wang M, Yuan J R, Qin L M, et al. TaCYP81D5, one member in a wheat cytochrome P 450 gene cluster, confers salinity tolerance via reactive oxygen species scavenging. Plant Biotechnology Journal, 2020, 18(3):791-804. |
| [31] | Kajino T, Yamaguchi M, Oshima Y, et al. KLU/CYP78A5, a cytochrome P450 monooxygenase identified via fox hunting, contributes to cuticle biosynthesis and improves various abiotic stress tolerances Frontiers in Plant Science, 2022, 13:904121. |
| [32] |
Zhang D, Yang H F, Wang X C, et al. Cytochrome P450 family member CYP96B5 hydroxylates alkanes to primary alcohols and is involved in rice leaf cuticular wax synthesis. New Phytologist, 2020, 225(5):2094-2107.
doi: 10.1111/nph.16267 pmid: 31618451 |
| [33] |
王家利, 刘冬成, 郭小丽, 等. 生长素合成途径的研究进展. 植物学报, 2012, 47(3):292-301.
doi: 10.3724/SP.J.1259.2012.00292 |
| [34] | Zhao Y, Hull A K, Gupta N R, et al. Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450s CYP79B2 and CYP79B3. Genes & Development, 2002, 16(23):3100-3112. |
| [35] | Tamiru M, Undan J R, Takagi H, et al. A cytochrome P450, OsDSS1, is involved in growth and drought stress responses in rice (Oryza sativa L.). Plant Molecular Biology, 2015, 88:85-99. |
| [36] | Kushiro T, Okamoto M, Nakabayashi K, et al. The Arabidopsis cytochrome P 450 CYP707A encodes ABA 8′‐hydroxylases: key enzymes in ABA catabolism. The EMBO Journal, 2004, 23(7):1647-1656. |
| [37] |
Sasaki E, Ogura T, Takei K, et al. Uniconazole, a cytochrome P450 inhibitor, inhibits trans-zeatin biosynthesis in Arabidopsis. Phytochemistry, 2013, 87:30-38.
doi: 10.1016/j.phytochem.2012.11.023 pmid: 23280040 |
| [1] | Teng Wen, Ye Fan, Zhou Zhou, Wang Yule, Liu Lijun. Effects of Wheat and Rapeseed Straw Returning on Yield and Quality of Rice under Salt Stress [J]. Crops, 2025, 41(5): 11-18. |
| [2] | Peng Binfeng, Lu Chusheng, Yin Yuanhong, Zhu Feifei, Ye Qunhuan, Pan Junfeng, Liu Yanzhuo, Hu Xiangyu, Hu Rui, Li Meijuan, Wang Xinyu, Liang Kaiming, Fu Youqiang. Physiological Mechanism of Ammonium-Nitrate Mixed Nutrition Promoting Rice Growth under High-Temperature Stress [J]. Crops, 2025, 41(5): 165-170. |
| [3] | Du Hanmeng, Chen Yuqiong, Liu Ruotong, Chen Yinglong, Dai Qigen, Zhang Hongcheng, Liao Ping. Effects of Chlormequat Chloride and Gypsum Application on Rice Yield and Lodging Risk under Salt Stress [J]. Crops, 2025, 41(5): 29-34. |
| [4] | Yang Linsheng, Xi Min, Tu Debao, Li Zhong, Zhou Yongjin, Xu Youzun, Sun Xueyuan, Wu Wenge. Assessment of Resource Input and Carbon, Nitrogen Footprint for Major Types of Rice in Yangtze River Delta Region [J]. Crops, 2025, 41(4): 150-156. |
| [5] | Deng Zhou, Gong Chenxu, He Yuxuan, Zeng Yongjun, Huang Shan. Effects of Combined Lime and Pig Manure Application on Grain Quality of Double-Cropping High-Quality Rice [J]. Crops, 2025, 41(4): 181-187. |
| [6] | He Xinchun, Du Hewei, Huang Min. Cloning and Expression Analysis of Maize ZmCaM1 Gene [J]. Crops, 2025, 41(4): 19-28. |
| [7] | Tao Zuhao, Wang Weiqin, Zheng Huabin, Xiang Jun, Tang Qiyuan. Effects of Water, Fertilizer and Chemical Regulation on Seedling Quality of Mechanized Casting Transplanting in Late Rice [J]. Crops, 2025, 41(4): 224-230. |
| [8] | Bu Qing, Yang Kaiqiang, Wang Tian, Shi Si, Xin Guochen, Chen Yong, Dong Zhaoxia, An Jing. Research Progress on the Use of Nanomaterials and the Alleviation of Abiotic Stresses in Crops [J]. Crops, 2025, 41(4): 9-18. |
| [9] | He Bing, Wang Xiaohang, Li Chao, Luo Liqiang, Zhang Qiang, Han Kangshun, Chen Dianyuan, Yan Guangbin, Liu Zhenjiao. Data Analysis of Approved Rice Varieties in Jilin Province from 1987 to 2022 [J]. Crops, 2025, 41(3): 16-22. |
| [10] | Cao Zhengnan, Zhao Zhendong, Hu Bo, Yu Han, Ning Xiaohai, Zhao Zeqiang, Cao Liyong. Effects of Nitrogen Fertilizer and Promoting Rot Bacteria Fertilizer on Decomposition Effect of Returning Rice Straw to Field and Yield in Cold Regions [J]. Crops, 2025, 41(3): 172-177. |
| [11] | Li Hu, Huang Qiuyao, Wu Zishuai, Liu Guanglin, Chen Chuanhua, Luo Qunchang, Zhu Qinan. Effects of Planting Density and Nitrogen Application Rate on Yield and Rice Quality of High-Quality Conventional Rice Guiyu 12 [J]. Crops, 2025, 41(3): 195-201. |
| [12] | Mao Shunxin, Xiao Wuwei, Zhang Zuolin, Huang Jiada, Wang Fei, Huang Jianliang, Peng Shaobing, Cui Kehui. Effects of Different Irrigation Patterns and Fertilizer Managements on the Growth of Axillary Buds and Yield Formation of Ratoon Season in Ratoon Rice [J]. Crops, 2025, 41(3): 92-101. |
| [13] | Chen Hu, Gao Yuan, Sun Jiameng, Yu Peng, Xiao Hongwu, Zhang Haitao. Identification and Bioinformatics Analysis of the Typical Thioredoxin (TRX) Gene Family in Barley [J]. Crops, 2025, 41(2): 66-73. |
| [14] | Zhao Fuyang, Ma Bo, Hu Jifang, Tan Kefei, Liu Chuanzeng, Yan Feng, Dong Yang, Hou Xiaomin, Li Qingquan, Han Yehui. Evaluation of Photoperiod Sensitivity of Japonica Rice in Cold Regions under Different Photoperiod Conditions [J]. Crops, 2025, 41(2): 135-140. |
| [15] | Ji Jinghong, Liu Shuangquan, Ma Xingzhu, Hao Xiaoyu, Zheng Yu, Zhao Yue, Wang Xiaojun, Kuang Enjun. Effects of Different Controlled-Release Urea on Agronomic Traits, Yield and Nitrogen Use Efficiency of Cold Region Rice [J]. Crops, 2025, 41(2): 149-154. |
|
||