Crops ›› 2025, Vol. 41 ›› Issue (6): 83-90.doi: 10.16035/j.issn.1001-7283.2025.06.011
Previous Articles Next Articles
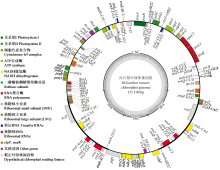
Chloroplast Genome Characteristics and Codon Bias Analysis of Anti-Broomrape Confectionery Sunflower Hybrid Jingkui 1408
Ma Jun1( ), Zhang Xianglei2, An Chengjie2, Yang Xuewen3, Zhang Bolin2, Zhang Wu2(
), Zhang Xianglei2, An Chengjie2, Yang Xuewen3, Zhang Bolin2, Zhang Wu2( ), Wang Pengdong4(
), Wang Pengdong4( )
)
- 1
Institute of Industrial Crops ,Heilongjiang Academy of Agricultural Sciences Harbin 150086, Heilongjiang, China
2College of Science ,Jiamusi University Jiamusi 154007, Heilongjiang, China
3College of Biology and Agriculture ,Jiamusi University Jiamusi 154007, Heilongjiang, China
4Research Institute of Economic Crops ,Shanxi Agricultural University, Taiyuan 030031 Shanxi, China
| [1] | 陈晓丹, 冯玉玺, 郭海洋, 等. 裸果木叶绿体基因组密码子使用偏性分析. 分子植物育种, 2023, 21(23):7715-7724. |
| [2] | 吴宪明, 吴松锋, 任大明, 等. 密码子偏性的分析方法及相关研究进展. 遗传, 2007, 29(4):420-426. |
| [3] | 赵耀, 刘汉梅, 顾勇, 等. 玉米waxy基因密码子偏好性分析. 玉米科学, 2008, 16(2):16-21. |
| [4] | 唐晓芬, 陈莉, 马玉韬. 密码子使用偏性量化方法研究综述. 基因组学与应用生物学, 2013, 32(5):660-666. |
| [5] | Sharp P M, Matassi G. Codon usage and genome volution. Current Opinion in Genetics & Development, 1994, 4(6):851-860. |
| [6] | Dierckxsens N, Mardulyn P, Smits G. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Research, 2017, 45(4):e18. |
| [7] | Tillich M, Lehwark P, Pellizzer T, et al. GeSeq-versatile and accurate annotation of organelle genomes. Nucleic Acids Research, 2017, 45(1):6-11. |
| [8] | 李冬梅, 吕复兵, 朱根发, 等. 文心兰叶绿体基因组密码子使用的相关分析. 广东农业科学, 2012, 39(10):61-65. |
| [9] | 王雪, 马军, 杨学文, 等. 栽培型向日葵叶绿体基因组密码子使用偏好性分析. 分子植物育种, 2024, 22(4):1096-1102. |
| [10] | 原晓龙, 李云琴, 王毅, 等. 西南桦叶绿体基因组密码子偏好性分析. 基因组学与应用生物学, 2020, 39(12):5758-5764. |
| [11] | 颜超, 徐钦, 李志, 等. 黎平瘤果茶的叶绿体基因组特征及系统发育分析. 分子植物育种,(2024-02-29)[2024-07-26]. https://link.cnki.net/urlid/46.1068.S.20240228.1659.008. |
| [12] |
杨国锋, 苏昆龙, 赵怡然, 等. 蒺藜苜蓿叶绿体密码子偏好性分析. 草业学报, 2015, 24(12):171-179.
doi: 10.11686/cyxb2015016 |
| [13] | 张文达, 赵逸伦, 王湘雨, 等. 再力花叶绿体基因组特征和系统发育分析. 分子植物育种,(2024-04-03)[2024-07-26]. https://link.cnki.net/urlid/46.1068.s.20240403.0936.006. |
| [14] | 崔雨丝, 韩骐泽, 张诗文, 等. 绣球叶绿体基因组特征及密码子偏好性分析. 分子植物育种,(2024-03-29)[2024-07-26]. https://link.cnki.net/urlid/46.1068.S.20240329.0938.007. |
| [15] | 李志芳, 陈丽玲, 罗淑洁, 等. 西南牡蒿叶绿体基因组特征及系统发育分析. 广西植物, 2024, 44(9):1732-1745. |
| [16] |
Hunt R C, Simhadri V L, Iandoli M, et al. Exposing synonymous mutations. Trends Genet, 2014, 30(7):308-321.
doi: 10.1016/j.tig.2014.04.006 pmid: 24954581 |
| [17] | 马楠, 叶奕含, 邹晨鑫, 等. 人参叶绿体全基因组特征及密码子偏好性分析. 分子植物育种,(2022-11-30)[2024-07-26]. https://link.cnki.net/urlid/46.1068.S.20221129.1638.028. |
| [18] |
余志伟, 陈垣, 郭凤霞, 等. 菘蓝叶绿体基因组密码子使用偏好性分析. 草地学报, 2024, 32(8):2374-2385.
doi: 10.11733/j.issn.1007-0435.2024.08.005 |
| [19] | 代国娜, 尚明越, 王嘉乐, 等. 百合属药用植物叶绿体基因组密码子偏好性及系统发育研究. 中草药, 2024, 55(11):3835-3844. |
| [20] | 黄思琦, 张麒功, 叶泽霖, 等. 5种柏科植物叶绿体基因组密码子偏好性分析. 福建农林大学学报, 2024, 53(2):214-220. |
| [21] | 魏亚楠, 龚明贵, 白娜, 等. 梁山慈竹叶绿体基因组密码子偏好性分析. 浙江农林大学学报, 2024, 41(4):696-705. |
| [22] | 张智, 张国帅, 张新可, 等. 硬尖神香草和欧神香草叶绿体基因组密码子偏好性分析. 分子植物育种,(2024-04-22)[2024-07-26]. https://link.cnki.net/urlid/46.1068.S.20240420.1355.002. |
| [23] | 谢刘义, 任璇, 应文博, 等. 栽培大麦和野生大麦叶绿体基因组密码子偏好性比较分析. 云南民族大学学报, 2024, 34(3):268-276. |
| [1] | Zhou Fei, Huang Xutang, Xie Pengyuan, Wang Jing, Liu Yan, Cui Jiawei, Tang Lili, Wang Wenjun. Construction of a TRV-Mediated VIGS System in Sunflower [J]. Crops, 2025, 41(6): 45-50. |
| [2] | Wang Heya, Luo Jingjing, Meng Ling, Ai Haifeng, Wang Bin, Li Huaisheng, Xu Jingpeng, Xu Xiangyang. Yield Sensitivity Analysis of Edible Sunflower Varieties in Ta'e Basin [J]. Crops, 2025, 41(3): 30-37. |
| [3] | Di Na, Zheng Xiqing, Wang Jing, Han Haijun, Li Na. Study on the Difference of Physiological Response of Sunflower to Broomrape Parasitism [J]. Crops, 2025, 41(2): 123-127. |
| [4] | Mu Jianguo, Wang Peng, Liu Yantao, Cui Jiawei, Chen Yanfang, Wan Sumei, Chen Guihong. Effects of Different Harvesting Periods on the Commerciality and Yield of Edible Sunflower [J]. Crops, 2024, 40(5): 146-151. |
| [5] | Liu Chunhui, Du Ruixia, Wang Yongxing, Yang Qinfang, Shan Feibiao, Xu Fuxun, Zhao Yuxin, Chen Shushu, Liu Wei. Relationship between Meteorological Factors and DUS Test Measurement Traits of Sunflower [J]. Crops, 2024, 40(3): 156-162. |
| [6] | Du Chao, Li Jun, Wang Gang, Wu Xuerui, Ren Zhiyuan, Zhang Junfeng, Bao Haizhu, Wen Aiqing. Effects of High Ridge Mulching Drip Irrigation on the Growth and Water Use of Sunflower in Moderate Saline-Alkali Land in Hetao Irrigation Region [J]. Crops, 2024, 40(1): 111-116. |
| [7] | Lü Zengshuai, Dong Hongye, Wang Peng, Duan Wei, Liu Shengli, Liu Yantao. Progress in Mechanism of Herbicide Resistance and Breeding of Sunflower [J]. Crops, 2024, 40(1): 16-22. |
| [8] | Wu Sheng, Duan Yu, Zhang Tingting, An Hao, Zhang Jun, Liang Junmei, Zhang Sheng. Relationships between Dry Matter Accumulation, Transport and Yield of Confectionary Sunflower and Response to Water and Nitrogen Interactions [J]. Crops, 2023, 39(6): 243-251. |
| [9] | Li Hepeng, Zhang Yunhua, Meng Qinglin, Ma Ligong, Yu Hongtao, Li Haiyan, Li Yichu, Liu Jia, Shi Fengmei, Yang Fan, Liu Liang. Screening of Inducers for Sunflower Sclerotinia sclerotiorum and Application of Hypersensitive Protein [J]. Crops, 2023, 39(6): 257-260. |
| [10] | Ling Yibo, Wang Binjie, Hu Yimin, Heinar·Madithermic mann, Chen Nianlai. Responses of Dry Matter Translocation and Yield Formation to Planting Density and Row Spacing of Sunflower [J]. Crops, 2023, 39(5): 197-203. |
| [11] | Yi Bing, Liu Jingang, Song Dianxiu, Wang Dexing, Zhao Mingzhu, Liu Xiaohong, Sun Enyu, Cui Liangji. Study on Land Productivity and Interspecific Competition of Sunflower and Millet Intercropping in Arid Areas [J]. Crops, 2023, 39(5): 219-223. |
| [12] | Zhu Kongyan, Han Shengcai, Zhao Rong, Wen Yujie, Hu Haochi, Qiao Yimin, Lu Jiafeng, Cao Kai, Xu Zhenghan, Bao Haizhu, Gao Julin. Isolation and Identification of Endophytes from Sunflower Seeds [J]. Crops, 2023, 39(5): 280-284. |
| [13] | Jia Xiuping, Mao Xuhui, Liang Gensheng, Liu Runping, Liu Feng, Wang Xingzhen. Analysis of Physiological and Biochemical Mechanism and Growth and Development Characteristics of Saline and Alkali Resistance in Sunflower [J]. Crops, 2022, 38(5): 146-152. |
| [14] | Shi Bixian, Tao Jianfei, Gao Yan, Xie Huihong, Abulimiti·Aierken , Cheng Pingshan, Maitituersun·Sadike , Sha Hong. Effects of Different Planting Densities on the Morphological Traits and Yields of Three Confectionery Sunflower Varieties [J]. Crops, 2022, 38(5): 195-200. |
| [15] | Zhou Fei. Bioinformatics and Expression Analysis of HaLACS7 Gene in Sunflower [J]. Crops, 2022, 38(3): 104-108. |
|
||