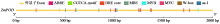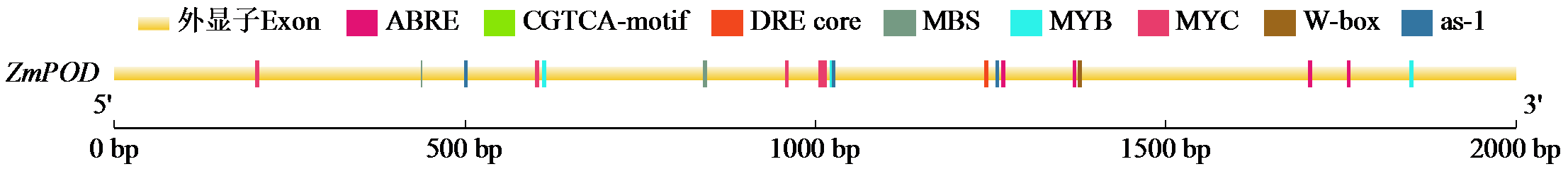Crops ›› 2025, Vol. 41 ›› Issue (6): 37-44.doi: 10.16035/j.issn.1001-7283.2025.06.005
Previous Articles Next Articles
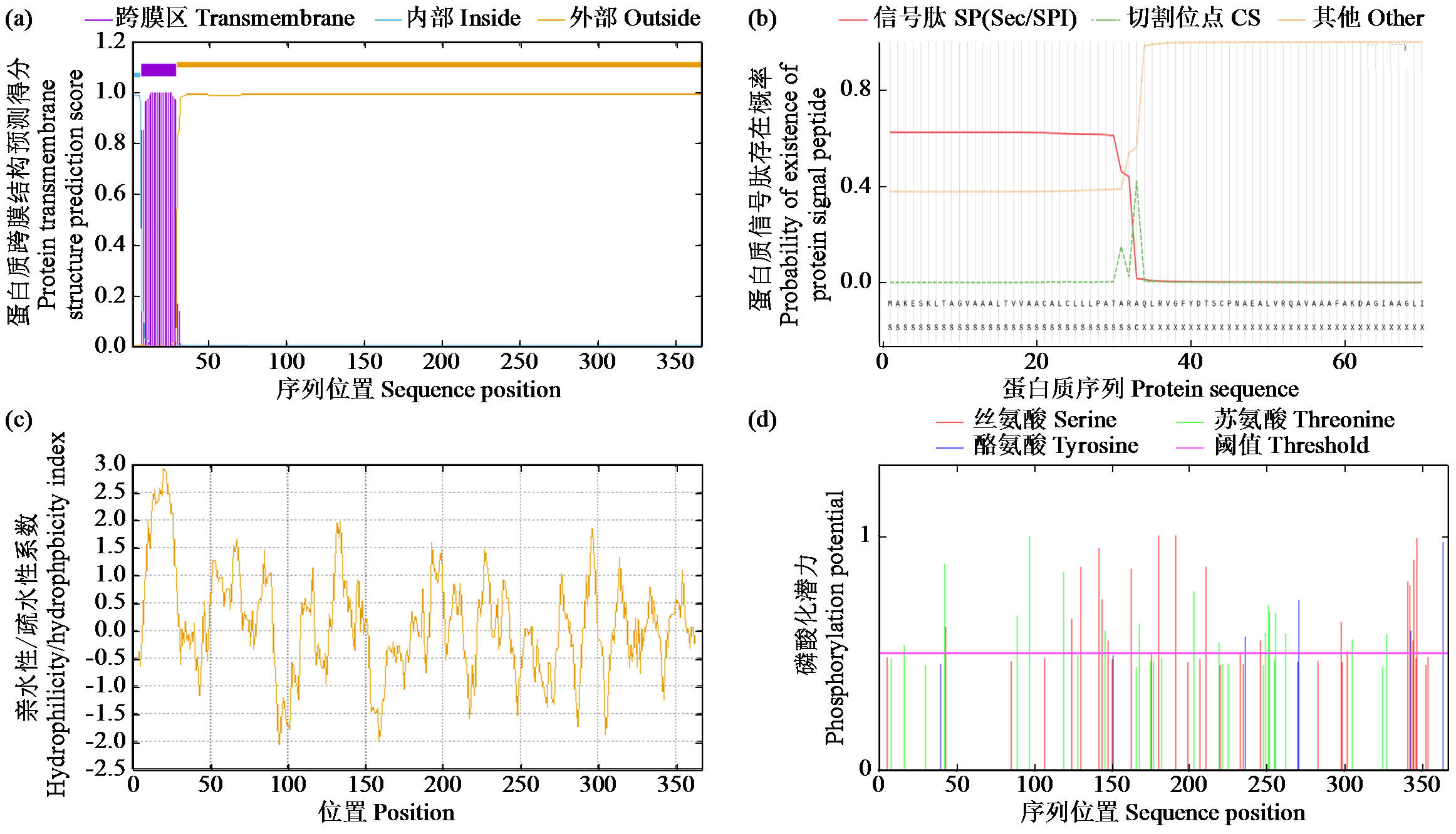
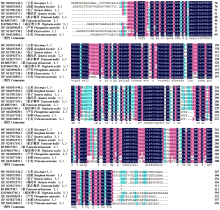
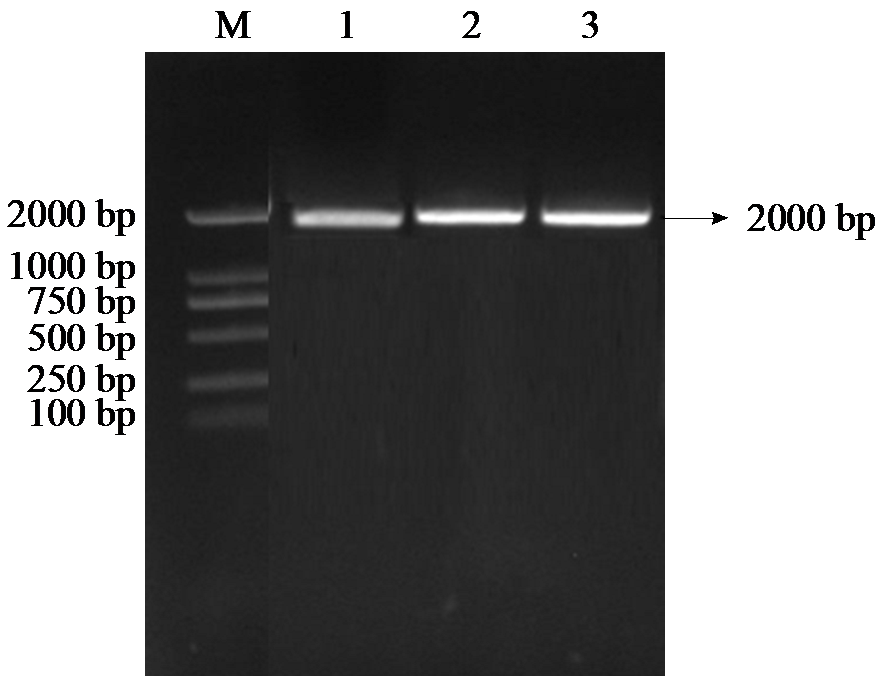
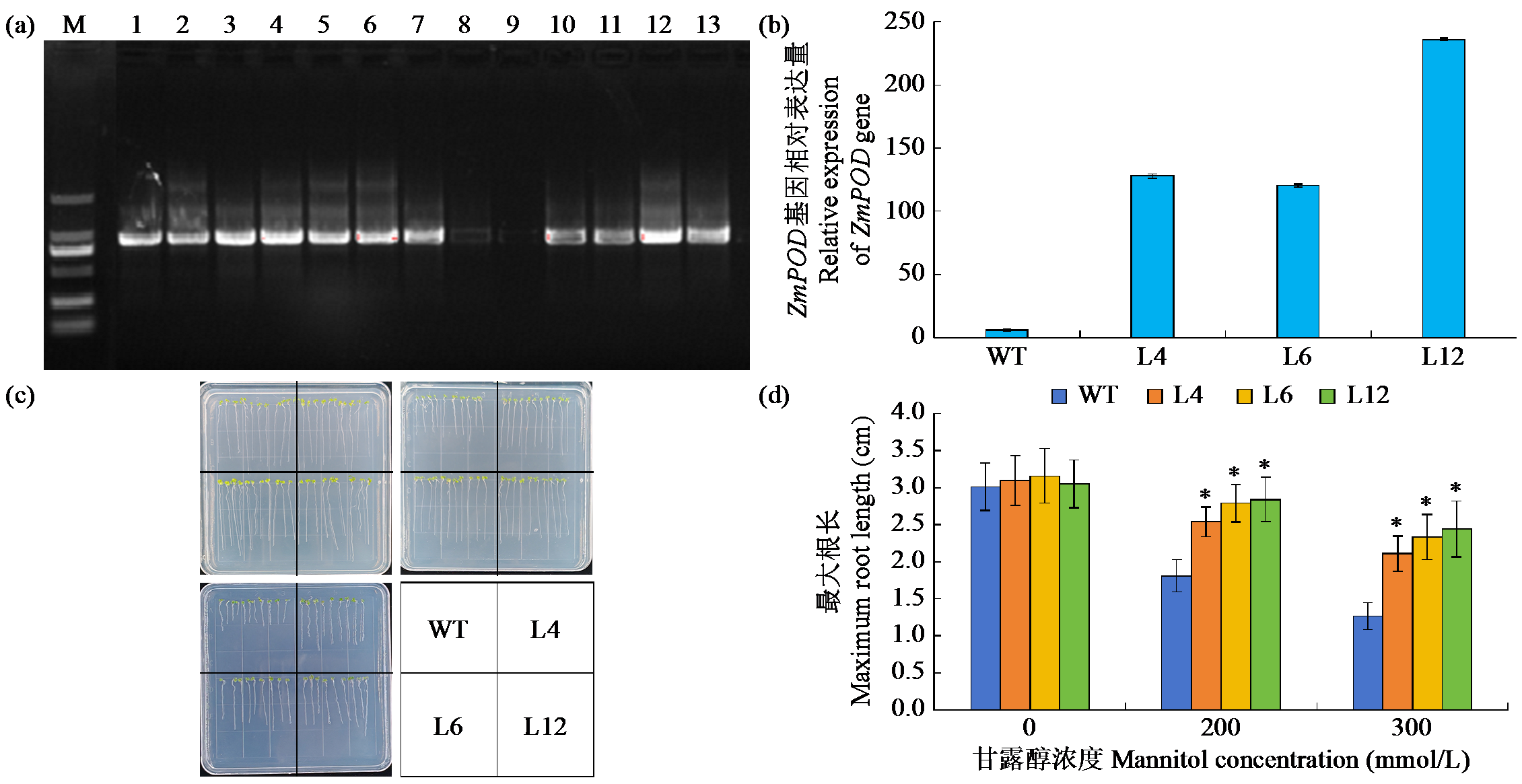
Cloning, Bioinformatics Analysis, and Functional Validation of ZmPOD Gene in Maize
Liu Songtao( ), Jiang Chao, Shi Hanbo, Yan Linan, Zhao Haichao, Lu Haibo, Li Hui, Huang Zhihong(
), Jiang Chao, Shi Hanbo, Yan Linan, Zhao Haichao, Lu Haibo, Li Hui, Huang Zhihong( )
)
Hebei North University / The Key Laboratory of Hebei Province Agricultural Products and Food Quality and Safety Analysis and Testing ,Zhangjiakou 075000 Hebei, China
| [1] | Aslam M, Maqbool M A, Cengiz R. Drought Stress in Maize (Zea mays L.). Switzerland:Springer, 2015. |
| [2] | 方缘, 张玉书, 米娜, 等. 干旱胁迫及补水对玉米生长发育和产量的影响. 玉米科学, 2018, 26(1):89-97. |
| [3] | 冯健英, 许洛, 王绍新, 等. 北方地区夏玉米主推品种的抗旱性研究. 安徽农业科学, 2015, 43(28):47-49. |
| [4] | 高志勇, 谢恒星, 李吉锋, 等. 玉米抗旱性鉴定指标及分子生物学机理研究. 陕西农业科学, 2017, 63(2):57-61. |
| [5] |
Miao Z Y, Han Z X, Zhang T, et al. A systems approach to a spatio-temporal understanding of the drought stress response in maize. Scientific Reports, 2017, 7:6950.
doi: 10.1038/s41598-017-04863-7 |
| [6] | Feller U, Vaseva I I. Extreme climatic events: impacts of drought and high temperature on physiological processes in agronomically important plants. Frontiers in Environmental Science, 2014, 2:39. |
| [7] | FAO. How to Feed the World in 2050. (2009-10-08)[2024-07-22]. https://www.fao.org/fileadmin/templates/wsfs/docs/expert_paper/How_to_Feed_the_World_in_2050.pdf. |
| [8] | 李跃. 不同基因型玉米自交系对干旱胁迫的生理特性分析. 杨凌:西北农林科技大学, 2014. |
| [9] |
Passardi F, Longet D, Penel C, et al. The class III peroxidase multigenic family in rice and its evolution in land plants. Phytochemistry, 2004, 65(13):1879-1893.
doi: 10.1016/j.phytochem.2004.06.023 pmid: 15279994 |
| [10] | Hiraga S, Sasaki K, Ito H, et al. A large family of class III plant peroxidases. Plant & Cell Physiology, 2001, 42(5):462-468. |
| [11] | Welinder K G. Plant peroxidases: structure function relationships. Geneva:University of Geneva, 1992. |
| [12] | Edwards S L, Raag R, Wariishi H, et al. Crystal structure of lignin peroxidase. Proceedings of the National Academy of Sciences of the United States of America, 1993, 90(2):750-754. |
| [13] |
Smulevich G, Jakopitsch C, Droghetti E, et al. Probing the structure and bifunctionality of catalase-peroxidase (KatG). Journal of Inorganic Biochemistry, 2006, 100(4):568-585.
pmid: 16516299 |
| [14] |
Passardi F, Bakalovic N, Teixeira F K, et al. Prokaryotic origins of the non-animal peroxidase superfamily and organelle-mediated transmission to eukaryotes. Genomics, 2007, 89(5):567-579.
pmid: 17355904 |
| [15] |
Skulachev V P. Cytochrome c in the apoptotic and antioxidant cascades. FEBS Letters, 1998, 423(3):275-280.
doi: 10.1016/s0014-5793(98)00061-1 pmid: 9515723 |
| [16] | Erman J E, Vitello L B. Yeast cytochrome c peroxidase: mechanistic studies via protein engineering. Biochimica et Biophysica Acta, 2002, 1597(2):193-220. |
| [17] |
Ruiz-Dueñas F J, Camarero S, Pérez-Boada M, et al. A new versatile peroxidase from Pleurotus. Biochemical Society Transactions, 2001, 29(2):116-122.
doi: 10.1042/bst0290116 |
| [18] |
Cosio C, Dunand C. Specific functions of individual class III peroxidase genes. Journal of Experimental Botany, 2009, 60(2):391-408.
doi: 10.1093/jxb/ern318 pmid: 19088338 |
| [19] |
Nair P M G, Chung I M. A mechanistic study on the toxic effect of copper oxide nanoparticles in soybean (Glycine max L.) root development and lignification of root cells. Biological Trace Element Research, 2014, 162(1/2/3):342-352.
doi: 10.1007/s12011-014-0106-5 |
| [20] |
Bindschedler L V, Dewdney J, Blee K A, et al. Peroxidase- dependent apoplastic oxidative burst in Arabidopsis required for pathogen resistance. The Plant Journal, 2006, 47(6):851-863.
doi: 10.1111/tpj.2006.47.issue-6 |
| [21] |
Fang Y J, You J, Xie K B, et al. Systematic sequence analysis and identification of tissue-specific or stress-responsive genes of NAC tran scription factor family in rice. Molecular Genetics and Genomics, 2008, 280(6):547-563.
doi: 10.1007/s00438-008-0386-6 |
| [22] | 蔡佳文. 泛素途径基因SorUBC和SorRma1在龙葵(Solanum nigrum L.)抗非生物胁迫中的作用研究. 哈尔滨:哈尔滨师范大学, 2017. |
| [23] | 吴梦露, 李鹏, 于文清, 等. 水稻防御酶与其抗病性关系研究进展. 分子植物育种, 2024, 22(7):2413-2420. |
| [24] | 徐建霞, 郑常祥. 玉米抗旱生理生化指标的研究进展. 农技服务, 2018, 35(4):33-35. |
| [1] | Wang Zhi, Zhou Wenli, Zhao Yao, Liu Zheng, Li Congfeng, Zhang Renhe. Effects of New Chemical Control Combinations on Photosynthetic Performance and Yield Improvement of Maize [J]. Crops, 2025, 41(6): 112-120. |
| [2] | Zhou Tingfang, Li Ran, Liu Qianqian, Zhang Ze, Wang Zhenhua, Ma Baoxin, Lu Ming, Zhang Lin, Han Yehui, Yang Bo, Li Mingshun, Zhang Degui, Weng Jianfeng, Yong Hongjun, Xu Jingyu, Han Jienan, Li Xinhai. Analysis of Salt Tolerance at Germination Stage of 118 Maize Hybrids in Northeast China [J]. Crops, 2025, 41(5): 1-10. |
| [3] | Yang Ke, Lai Shangkun, Jin Qian, Wang Weijun, Ding Yidong, Liu Xiaofei. Study on the Mining and Transcriptional Regulation Mechanism of Genes Related to Linoleic Acid Synthesis in Maize [J]. Crops, 2025, 41(5): 102-112. |
| [4] | Wang Changliang, Yan Lihui, Chang Jianzhi, Zhang Guohe, Wang Jing, Wang Fenxia, Huang Lihua. Breeding and Innovative Idea of Maize New Variety Xundan 168 [J]. Crops, 2025, 41(5): 142-146. |
| [5] | Sun Xianyin, Zhang Jibo, Lü Guangde, Qi Xiaolei, Sun Yingying, Mi Yong, Mu Qiuhuan, Yin Xundong, Wang Ruixia, Qian Zhaoguo, Gao Minggang. Comparison of High and Stable Yield Characteristics of Different Genotypes of Wheat under Dryland and Supplemental Irrigation Conditions [J]. Crops, 2025, 41(4): 104-110. |
| [6] | Li Zhehao, Ji Miyuan, Lü Meng, Ming Bo, Li Shaokun, Zhang Haiyan, Xie Ruizhi. Effects of Different Nitrogen Fertilizer Management Strategies on the Root and Shoot Development of Spring Maize under Long-Term Positioning Conditions [J]. Crops, 2025, 41(4): 135-141. |
| [7] | Wang Xingya, Chen Yuhan, Zhang Mengwen, Sun Linlin, Chen Lirong, Guo Yuqiu, Gong Kuijie. The Effects of ABA Application at Different Stages on Maize Grain Filling and Dehydration [J]. Crops, 2025, 41(4): 173-180. |
| [8] | He Xinchun, Du Hewei, Huang Min. Cloning and Expression Analysis of Maize ZmCaM1 Gene [J]. Crops, 2025, 41(4): 19-28. |
| [9] | Zhang Zhihan, Yao Jie, Zhang Zhantian, Bian Fuhua, Zhang Ziran, Chen Ping, Chen Haining, Liu Baoyou. Effects of 5-Aminolevulinic Acid on Cucumber Seedling Growth and Soil Enzyme Activities under Drought Stress [J]. Crops, 2025, 41(4): 245-250. |
| [10] | Shi Yaxing, Liu Junling, Zhu Guichuan, He Zhongyou, Liu Hui, Fan Yanli, Xu Li, Lu Baishan, Zhao Jiuran, Luo Meijie. Cloning of the du1 Novel Allelic Variant and the Development of Its Molecular Markers [J]. Crops, 2025, 41(4): 41-48. |
| [11] | Hou Yue, Wang Hongliang, Li Jie, Li Chunjie, Chen Fanjun. Research Advances on the Effects of Cereal/Legume Forage Intercropping on Forage Quality and Nitrogen Uptake [J]. Crops, 2025, 41(3): 1-10. |
| [12] | Jin Zihao, Zhao Wenqing, Wang Fang, Wang Wei, Peng Yunling, Chang Fangguo. Effects of Three Exogenous Plant Growth Regulators on the Cold Tolerance of Maize Seedlings [J]. Crops, 2025, 41(3): 125-132. |
| [13] | Hou Xiaomin, Shen Huibo, Dong Shoukun, Yan Feng, Dong Yang, Zhao Fuyang, Li Qingquan, Zuo Yuetao. Physiological Effects of Mepiquat Chloride on Alleviating Drought Stress in Soybean Seedling Leaves [J]. Crops, 2025, 41(3): 133-140. |
| [14] | Wang Yi, Ren Yongfu, Zhang Zhengpeng, Ding Defang, Zhang Jing, Liu Yihong, Sun Duoxin, Chen Guangrong. The Effects of Different Covering Materials on Soil Environment and Maize Yield in Hexi Irrigation Area [J]. Crops, 2025, 41(3): 149-155. |
| [15] | Hou Nan, Wu Fengjie, Qi Xiangkun, Wang Yufeng, Yang Kejun, Fu Jian. Effects of Different Nitrogen Application Levels on Carbon Metabolism of Waxy Maize during Filling Period in Black Soil Area [J]. Crops, 2025, 41(3): 178-184. |
|
||