Crops ›› 2023, Vol. 39 ›› Issue (4): 31-37.doi: 10.16035/j.issn.1001-7283.2023.04.005
Previous Articles Next Articles
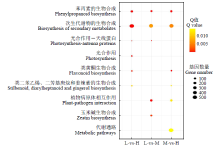
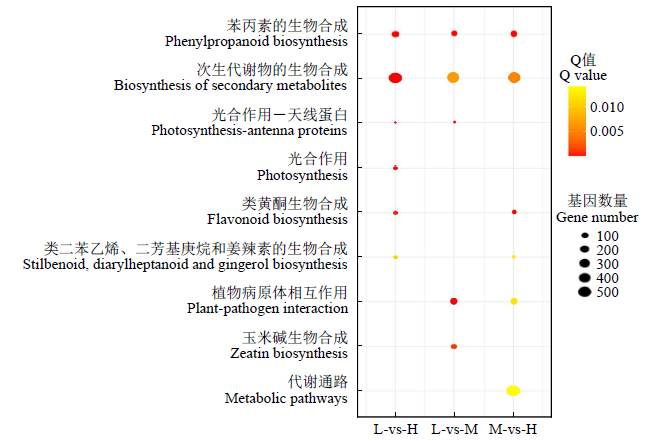
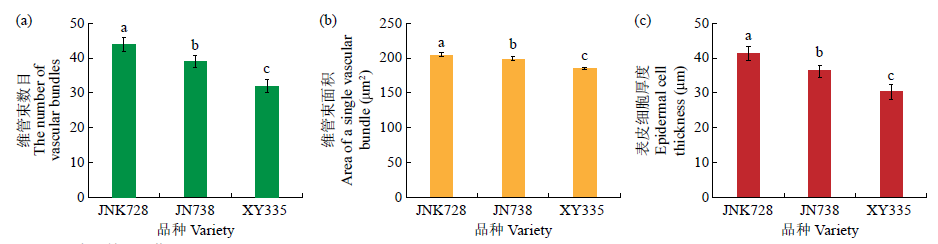
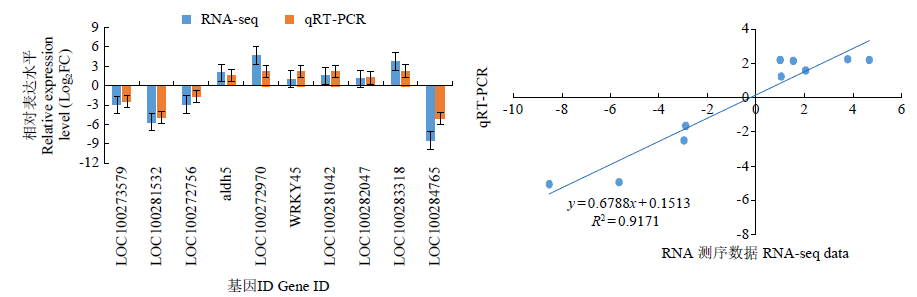
Transcriptomic Analysis to Reveal Lodging Resistance Genes and Metabolism Pathways in Maize (Zea mays L.)
Liu Songtao1( ), Tian Zaimin1, Liu Zigang1, Gao Zhijia2, Zhang Jing2, He Donggang3, Huang Zhihong1, Lan Xin1
), Tian Zaimin1, Liu Zigang1, Gao Zhijia2, Zhang Jing2, He Donggang3, Huang Zhihong1, Lan Xin1
- 1Hebei North University/Key Laboratory of Hebei Province Agricultural Products Food Quality and Safety Analysis and Testing, Zhangjiakou 075000, Hebei, China
2Hebei Universe Agricultural Science and Technology Co., Ltd., Zhangjiakou 075000, Hebei, China
3Hebei Zhaoyu Seed Industry Group Co., Ltd., Shijiazhuang 050000, Hebei, China
| [1] | 王聪, 马青美, 郭新梅, 等. 玉米茎秆抗倒伏相关性状的遗传效应分析. 玉米科学, 2021, 29(4):9-17. |
| [2] | 丰光, 黄长玲, 邢锦丰. 玉米抗倒伏的研究进展. 作物杂志, 2008(4):12-14. |
| [3] |
王夏青, 宋伟, 张如养, 等. 玉米茎秆抗倒伏遗传的研究进展. 中国农业科学, 2021, 54(11):2261-2272.
doi: 10.3864/j.issn.0578-1752.2021.11.002 |
| [4] | 马天军. 玉米倒伏种类及防治对策研究. 新农业, 2021(21):14. |
| [5] | 李妍妍, 景希强, 丰光, 等. 玉米倒伏的主要相关因素研究进展. 辽宁农业科学, 2013(4):47-51. |
| [6] | 徐天成, 周富亮, 黄联润, 等. 玉米抗倒伏性相关研究进展. 安徽农学通报, 2019, 25(11):29-32. |
| [7] |
Cloninger FR, Zuber MS. Methods for evaluating stalk quality in corn. Phytopathology, 1970, 60:295-300.
doi: 10.1094/Phyto-60-295 |
| [8] | 冯素伟, 姜小苓, 胡铁柱, 等. 不同小麦品种茎秆显微结构与抗倒强度关系研究. 中国农学通报, 2012, 28(36):57-62. |
| [9] |
LivakKJ, SchmittgenTD. Analysis of relative gene expression data using real-time quantitative PCR andthe 2-∆∆CT method. Methods, 2001, 25:402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [10] | 殷静. 大豆茎倒伏突变体基因定位、转录组测序及2个木质素合成基因的克隆研究.南京:南京农业大学, 2013. |
| [11] | 杨硕, 郑云霄, 黄亚群, 等. 不同玉米自交系茎秆上部维管束数目的差异分析. 植物遗传资源学报, 2021, 22(2):466-475. |
| [12] | 张巫军, 段秀建, 姚雄, 等. 烯效唑对遮阴下重穗型水稻渝香203茎秆形态结构和抗倒伏性的影响. 四川农业大学学报, 2019, 37(6):755-761. |
| [13] | 胡丹. 甜荞抗倒伏相关性状的遗传分析及木质素合成特征. 重庆:西南大学, 2016. |
| [14] | 刘丹. 木质素相关基因表达调控对油菜木质素合成及抗菌核病和抗倒伏性的影响. 北京: 中国农业科学院, 2008. |
| [1] | Zhou Wenli, Hao Miaoyi, Zhang Renhe. Effects of Nitrogen Fertilizer on Maize Root Growth and Nitrogen Metabolism under High-Density Planting [J]. Crops, 2026, 42(1): 125-132. |
| [2] | Ma Xiaoming, Qi Xiangkun, Tan Xue, Shi Mengyu, Wang Yufeng, Fu Jian, Yang Kejun. Effects of No-tillage with Straw Mulching on Soil Aggregate Stability and Maize Yield in Semi-Arid Region [J]. Crops, 2026, 42(1): 152-159. |
| [3] | Wang Zhi, Zhou Wenli, Zhao Yao, Liu Zheng, Li Congfeng, Zhang Renhe. Effects of New Chemical Control Combinations on Photosynthetic Performance and Yield Improvement of Maize [J]. Crops, 2025, 41(6): 112-120. |
| [4] | Liu Songtao, Jiang Chao, Shi Hanbo, Yan Linan, Zhao Haichao, Lu Haibo, Li Hui, Huang Zhihong. Cloning, Bioinformatics Analysis, and Functional Validation of ZmPOD Gene in Maize [J]. Crops, 2025, 41(6): 37-44. |
| [5] | Zhou Tingfang, Li Ran, Liu Qianqian, Zhang Ze, Wang Zhenhua, Ma Baoxin, Lu Ming, Zhang Lin, Han Yehui, Yang Bo, Li Mingshun, Zhang Degui, Weng Jianfeng, Yong Hongjun, Xu Jingyu, Han Jienan, Li Xinhai. Analysis of Salt Tolerance at Germination Stage of 118 Maize Hybrids in Northeast China [J]. Crops, 2025, 41(5): 1-10. |
| [6] | Yang Ke, Lai Shangkun, Jin Qian, Wang Weijun, Ding Yidong, Liu Xiaofei. Study on the Mining and Transcriptional Regulation Mechanism of Genes Related to Linoleic Acid Synthesis in Maize [J]. Crops, 2025, 41(5): 102-112. |
| [7] | Wang Changliang, Yan Lihui, Chang Jianzhi, Zhang Guohe, Wang Jing, Wang Fenxia, Huang Lihua. Breeding and Innovative Idea of Maize New Variety Xundan 168 [J]. Crops, 2025, 41(5): 142-146. |
| [8] | Li Zhehao, Ji Miyuan, Lü Meng, Ming Bo, Li Shaokun, Zhang Haiyan, Xie Ruizhi. Effects of Different Nitrogen Fertilizer Management Strategies on the Root and Shoot Development of Spring Maize under Long-Term Positioning Conditions [J]. Crops, 2025, 41(4): 135-141. |
| [9] | Wang Xingya, Chen Yuhan, Zhang Mengwen, Sun Linlin, Chen Lirong, Guo Yuqiu, Gong Kuijie. The Effects of ABA Application at Different Stages on Maize Grain Filling and Dehydration [J]. Crops, 2025, 41(4): 173-180. |
| [10] | He Xinchun, Du Hewei, Huang Min. Cloning and Expression Analysis of Maize ZmCaM1 Gene [J]. Crops, 2025, 41(4): 19-28. |
| [11] | Shi Yaxing, Liu Junling, Zhu Guichuan, He Zhongyou, Liu Hui, Fan Yanli, Xu Li, Lu Baishan, Zhao Jiuran, Luo Meijie. Cloning of the du1 Novel Allelic Variant and the Development of Its Molecular Markers [J]. Crops, 2025, 41(4): 41-48. |
| [12] | Hou Yue, Wang Hongliang, Li Jie, Li Chunjie, Chen Fanjun. Research Advances on the Effects of Cereal/Legume Forage Intercropping on Forage Quality and Nitrogen Uptake [J]. Crops, 2025, 41(3): 1-10. |
| [13] | Jin Zihao, Zhao Wenqing, Wang Fang, Wang Wei, Peng Yunling, Chang Fangguo. Effects of Three Exogenous Plant Growth Regulators on the Cold Tolerance of Maize Seedlings [J]. Crops, 2025, 41(3): 125-132. |
| [14] | Wang Yi, Ren Yongfu, Zhang Zhengpeng, Ding Defang, Zhang Jing, Liu Yihong, Sun Duoxin, Chen Guangrong. The Effects of Different Covering Materials on Soil Environment and Maize Yield in Hexi Irrigation Area [J]. Crops, 2025, 41(3): 149-155. |
| [15] | Hou Nan, Wu Fengjie, Qi Xiangkun, Wang Yufeng, Yang Kejun, Fu Jian. Effects of Different Nitrogen Application Levels on Carbon Metabolism of Waxy Maize during Filling Period in Black Soil Area [J]. Crops, 2025, 41(3): 178-184. |
|
||